ezTag is a web-based concept tagging tool that allows users to manually annotate text with biomedical concepts, use annotated data to train models, and use trained models to tag text automatically. Because ezTag is interactive, the automatically tagged text can then be refined manually to create new annotated data for training an improved model.
In ezTag, users can upload documents in BioC format, including PubMed abstracts and PubMed Central full-text articles. Biomedical concepts (biomedical named entities and their concept IDs) can then be annotated with one of several automated tools:
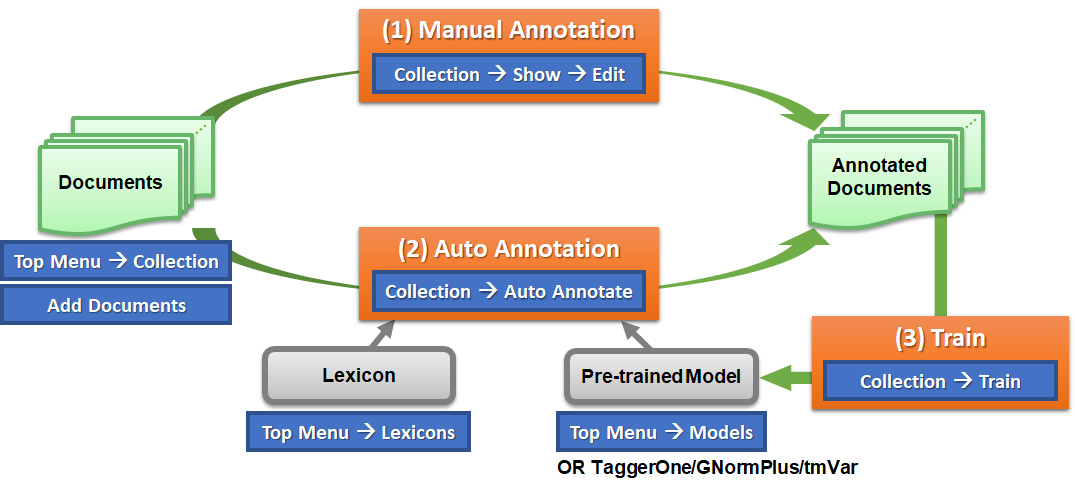
Basic workflow (without interactive learning):

- The blue boxes in the figure indicate the steps users should follow to navigate to the specified function.
PLEASE READ THE TUTORIAL FOR MORE DETAILS OR GO TO COLLECTIONS TO START ANNOTATING.
D. Kwon, S. Kim, C.-H. Wei, R. Leaman and Z. Lu, ezTag: tagging biomedical concepts via interactive learning, Nucleic Acids Research, 46, W523-W529, 2018.
Any entity type
ezTag supports a wide variety of entity types, not just common types (gene, chemical, disease names and sequence variants).
Full-text articles
The ezTag interface supports any document in BioC format, including both PubMed abstracts and PubMed Central full-text articles.
Custom lexicons
ezTag allows users to upload their own dictionaries for annotation with the string match algorithm and training our tagging module, TaggerOne.
Interactive learning
ezTag allows users to revise annotations made automatically rather than annotate completely manually, which may lead to better productivity.
Manual annotation
ezTag provides a user-friendly interface for manual annotation.
Programmatic APIs
Trained tagging models are also accessible through RESTful APIs for usage outside ezTag or large-scale application.